 
Proteins can indeed be represented in multiple, complementary ways, for example as sequences 22, 23, residue graphs 24, 25, 26, 27, atomic density maps 28, 29, 30, 31, 32, 33, 34, atomic point clouds 35 or molecular surfaces 36, 37, each capturing different functionally relevant features. Adapting the deep learning approach to protein structures requires defining an appropriate representation for proteins. 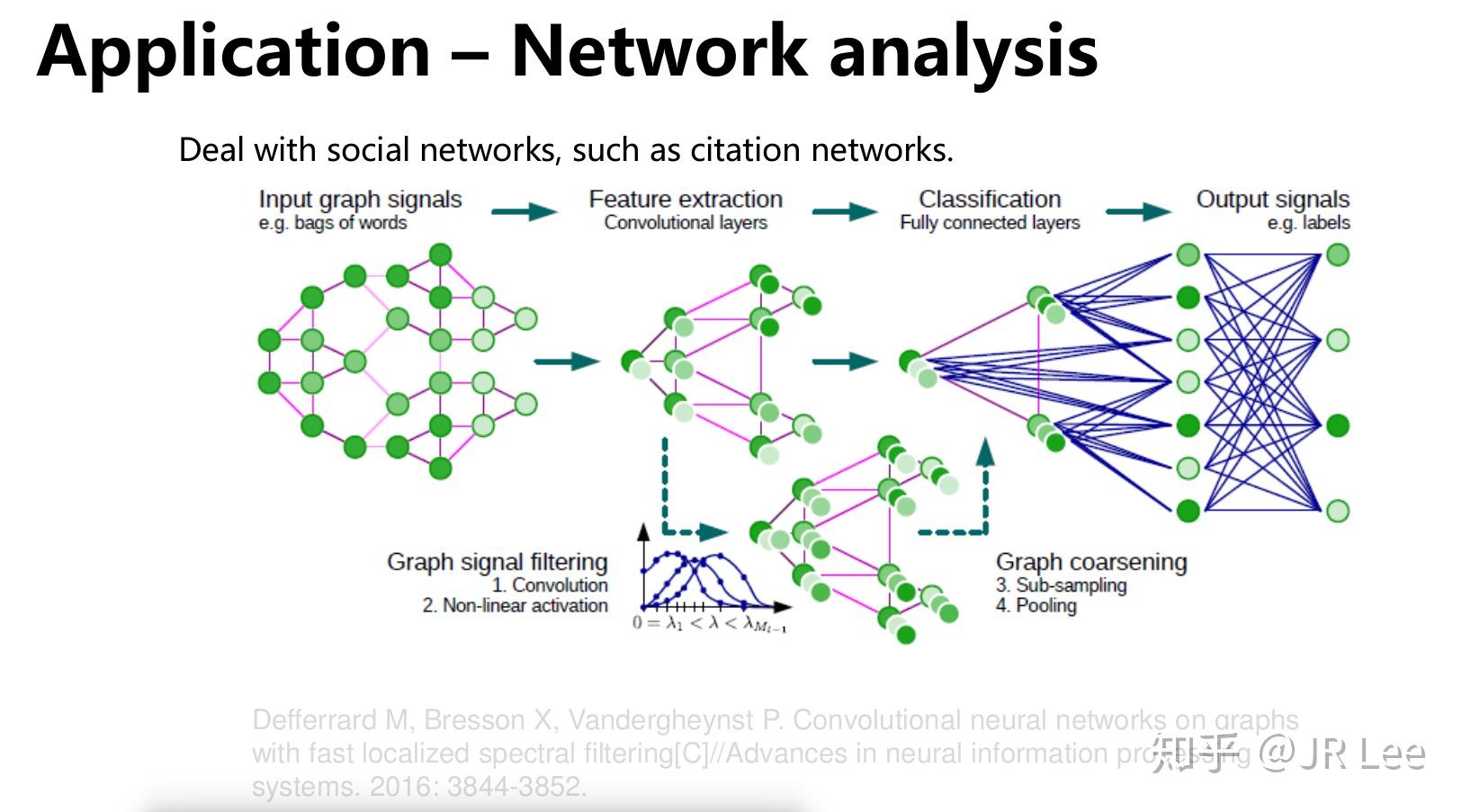
Indeed, deep learning models can learn the data features and their invariances directly by backpropagation, and generalize well despite a large number of parameters. 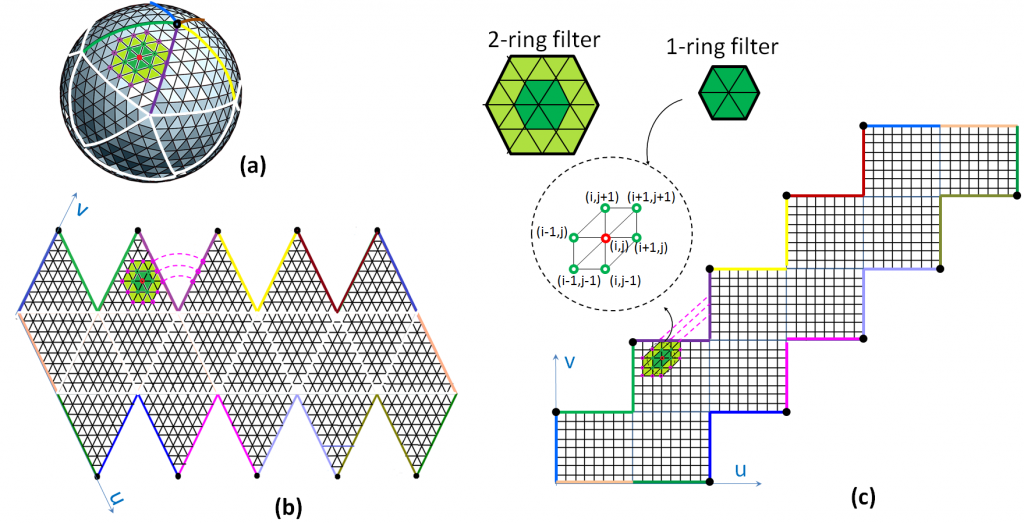
Despite over 50 years of experimental structural determination, new function-determining motifs are still being discovered 21.Įnd-to-end differentiable models, that is, deep learning, can potentially overcome the limitations of both approaches. Examples of such function-bearing motifs include Zinc fingers that are signatures of DNA or RNA binding sites 19, or PPI hotspot ‘O-rings’ 20: namely, exposed hydrophobic/aromatic amino acids surrounded by polar/charged ones. Machine learning models are, however, limited by the expressiveness of the features used, as these cannot capture the spatio-chemical arrangements of atoms or amino acids characterizing function-bearing motifs. Reasoning on mathematically defined features offers three advantages: (1) ability to generalize to proteins with no similarity to any of the train set proteins, (2) high sequence sensitivity, that is, ability to output distinct predictions for highly similar protein sequences and (3) fast inference speed. Then, the target property is predicted using a machine learning model for tabular data such as random forest or gradient boosting. For each amino acid of a query protein, various features of geometrical (for example, secondary structure, solvent accessibility, molecular surface curvature), physico-chemical (for example, hydrophobicity, polarity, electrostatic potential) and evolutionary (for example, conservation, position–weight matrices, coevolution) nature are calculated. This hampers our ability to both define and recognize such motifs using conventional comparative approaches.Īn alternative to comparative modeling is feature-based machine learning 12, 13, 14, 15, 16, 17, 18. Put differently, the invariances in both sequence and conformation spaces of such function-determining structural motifs are in general motif-dependent and therefore unknown. On the other hand, some protein–protein interactions (PPIs) are mainly driven by few ‘hotspot’ residues: mutations and/or conformational changes of the other interface residues preserve the interaction. On the one hand, the B cell epitopes (BCEs) of viral proteins frequently undergo antigenic drift, that is, the abolition of recognition by antibodies after only one or few mutations. Second, functional sites are variably preserved throughout evolution. First and foremost, its coverage is limited, as the pool of experimentally characterized protein folds or structural motifs is small. Comparative modeling has several shortcomings. The most accurate functional site prediction method is comparative modeling 5, 6, 7, 8, 9, 10, 11, 12, 13: given a query protein, similar proteins with known functional sites are searched for and their sites are mapped onto the query structure. ĭespite recent progresses in experimental 1 and AI-based 2, 3 protein structure determination, there remains a gap between structure and function 4. A webserver for ScanNet is available from. Overall, ScanNet is a versatile, powerful and interpretable model suitable for functional site prediction tasks. Finally, we predict epitopes of the SARS-CoV-2 spike protein, validating known antigenic regions and predicting previously uncharacterized ones. We train ScanNet for detecting protein–protein and protein–antibody binding sites, demonstrate its accuracy-including for unseen protein folds-and interpret the filters learned. ScanNet builds representations of atoms and amino acids based on the spatio-chemical arrangement of their neighbors. Here, we introduce ScanNet, an end-to-end, interpretable geometric deep learning model that learns features directly from 3D structures. They are, respectively, limited by the expressivity of the handcrafted features and the availability of similar proteins. Currently, two classes of methods prevail: machine learning models built on top of handcrafted features and comparative modeling. Predicting the functional sites of a protein from its structure, such as the binding sites of small molecules, other proteins or antibodies, sheds light on its function in vivo. 
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
